How to Read a Column From Csv File in R

Welcome! If you desire to offset diving into data science and statistics, and so data frames, CSV files, and R will be essential tools for you lot. Let'due south encounter how you can apply their amazing capabilities.
In this article, you will larn:
- What CSV files are and what they are used for.
- How to create CSV files using Google Sheets.
- How to read CSV files in R.
- What Data Frames are and what they are used for.
- How to access the elements of a data frame.
- How to alter a data frame.
- How to add and delete rows and columns.
Nosotros will utilize RStudio, an open-source IDE (Integrated Development Environment) to run the examples.
Let's begin! ✨
🔹 Introduction to CSV Files
CSV (Comma-separated Values) files tin be considered ane of the building blocks of information analysis because they are used to shop data represented in the form of a table.
In this file, values are separated by commas to represent the unlike columns of the tabular array, similar in this example:
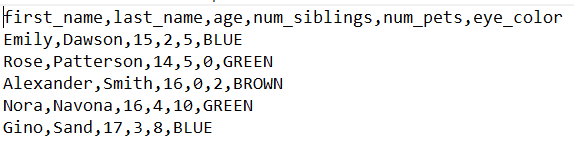
We will generate this file using Google Sheets.
🔸 How to Create a CSV File Using Google Sheets
Let's create your showtime CSV file using Google Sheets.
Step 1: Get to the Google Sheets Website and click on "Go to Google Sheets":
💡 Tip: You tin admission Google Sheets by clicking on the push button located at the top-correct edge of Google'south Home Page:
If nosotros zoom in, we see the "Sheets" button:
💡 Tip: To use Google Sheets, you need to accept a Gmail account. Alternatively, you can create a CSV file using MS Excel or another spreadsheet editor.
Y'all will see this panel:
Pace 2: Create a blank spreadsheet by clicking on the "+" button.
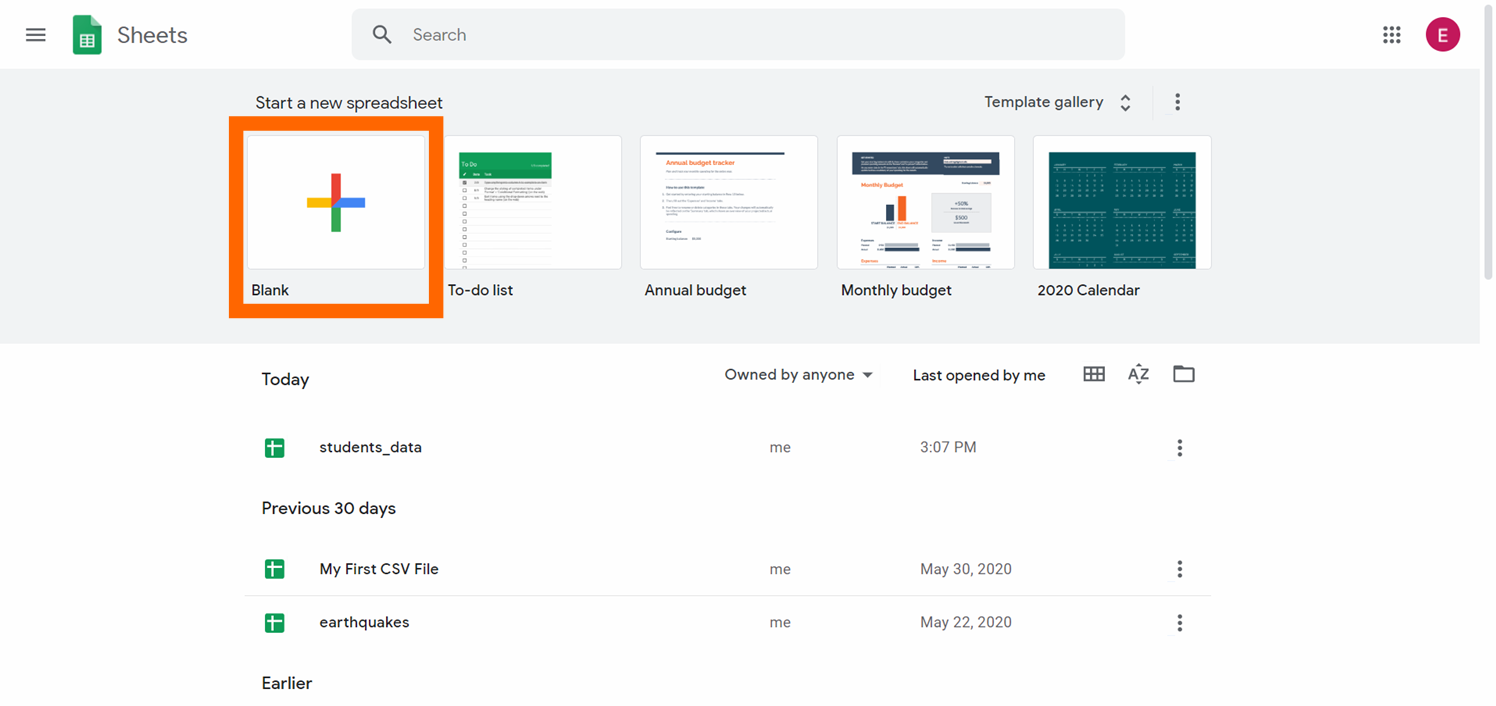
At present yous have a new empty spreadsheet:
Step 3: Change the name of the spreadsheet to students_data. We volition need to employ the proper name of the file to piece of work with data frames. Write the new name and click enter to confirm the change.
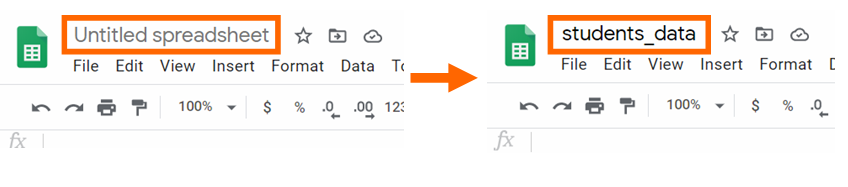
Step 4: In the first row of the spreadsheet, write the titles of the columns.
When you import a CSV file in R, the titles of the columns are called variables. Nosotros will define six variables: first_name, last_name, age, num_siblings, num_pets, and eye_color, as you can see right hither beneath:
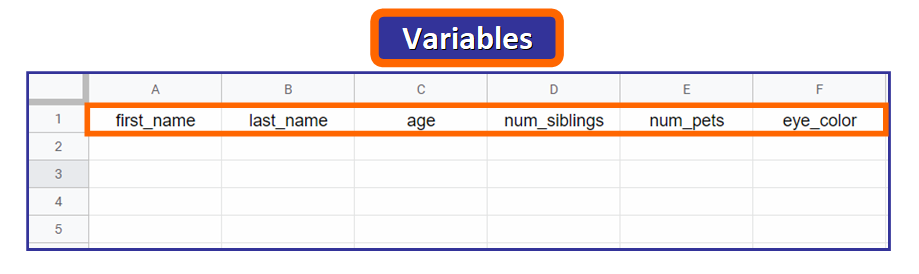

💡 Tip: Notice that the names are written in lowercase and words are separated with an underscore. This is not mandatory, merely since you will demand to access these names in R, it's very mutual to use this format.
Step 5: Enter the data for each one of the columns.
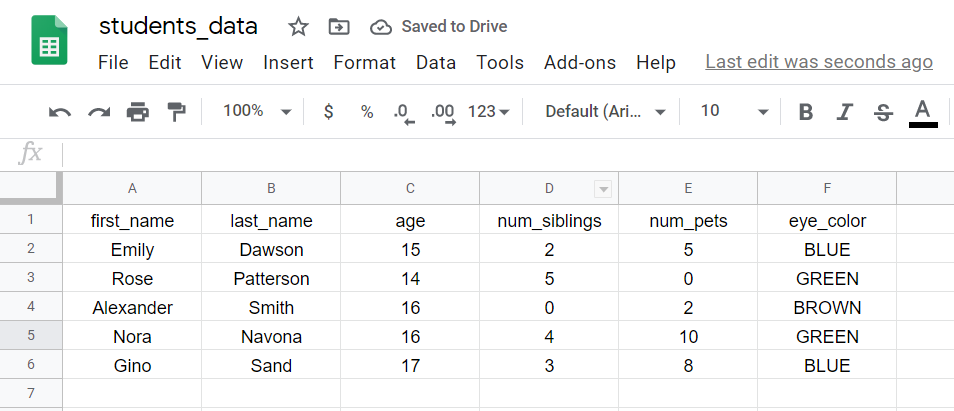
When you read the file in R, each row is chosen an observation, and it corresponds to information taken from an individual, animal, object, or entity that we collected data from.
In this case, each row corresponds to the data of a educatee:
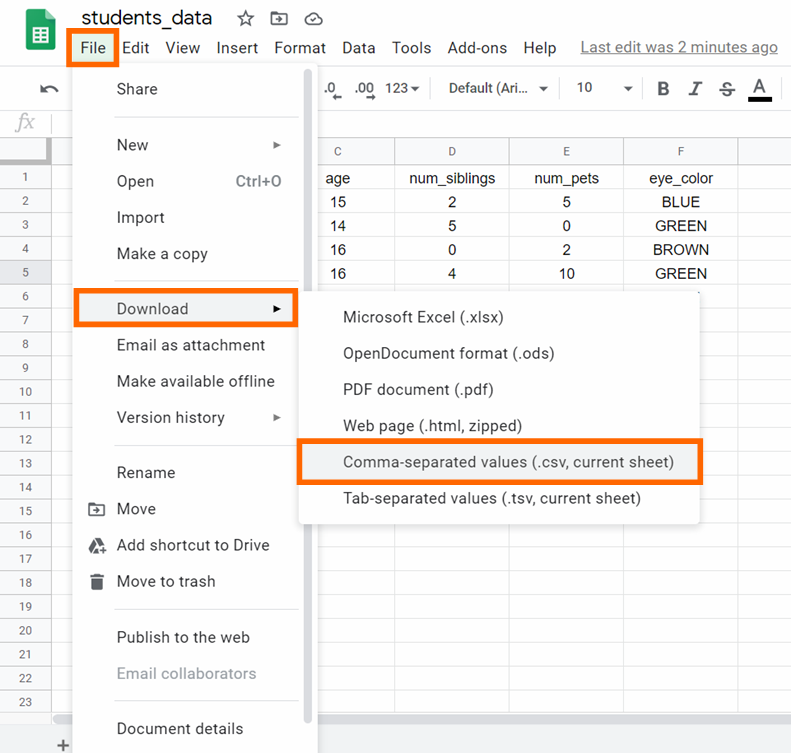
Stride 6: Download the CSV file by clicking on File -> Download -> Comma-separated values, equally you tin can see below:
Step 7: Rename the file CSV file. You volition need to remove "Sheet1" from the default name because Google Sheet volition automatically add this to the name of the file.

Great work! At present you accept your CSV file and it's time to start working with it in R.
🔹 How to Read a CSV file in R
In RStudio, the first step before reading a CSV file is making certain that your current working directory is the directory where the CSV file is located.
💡 Tip: If this is not the case, you lot will demand to utilise the total path to the file.
Change Current Working Directory
You lot can change your current working directory in this panel:
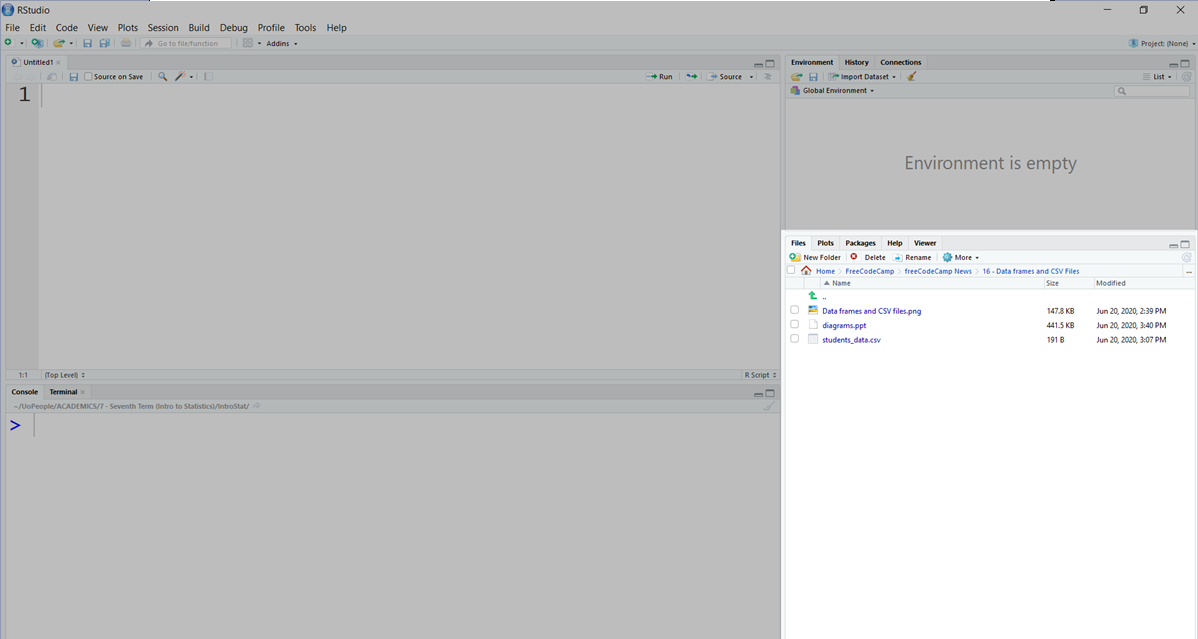
If we zoom in, y'all can see the current path (1) and select the new one by clicking on the ellipsis (...) push to the right (2):
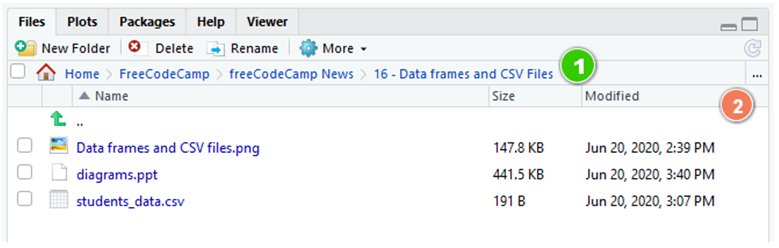
💡 Tip: You can also check your current working directory with getwd() in the interactive console.
And so, click "More than" and "Gear up Equally Working Directory".
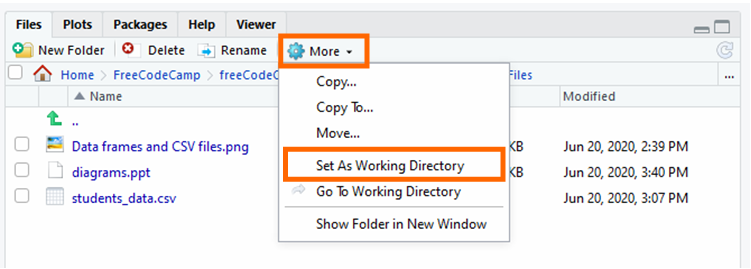
Read the CSV File
Once you take your electric current working directory set up, yous can read the CSV file with this command:

In R lawmaking, nosotros take this:
> students_data <- read.csv("students_data.csv") 💡 Tip: Nosotros assign information technology to the variable students_data to admission the data of the CSV file with this variable. In R, we can separate words using dots ., underscores _, UpperCamelCase, or lowerCamelCase.
After running this command, you lot will see this in the top right panel:
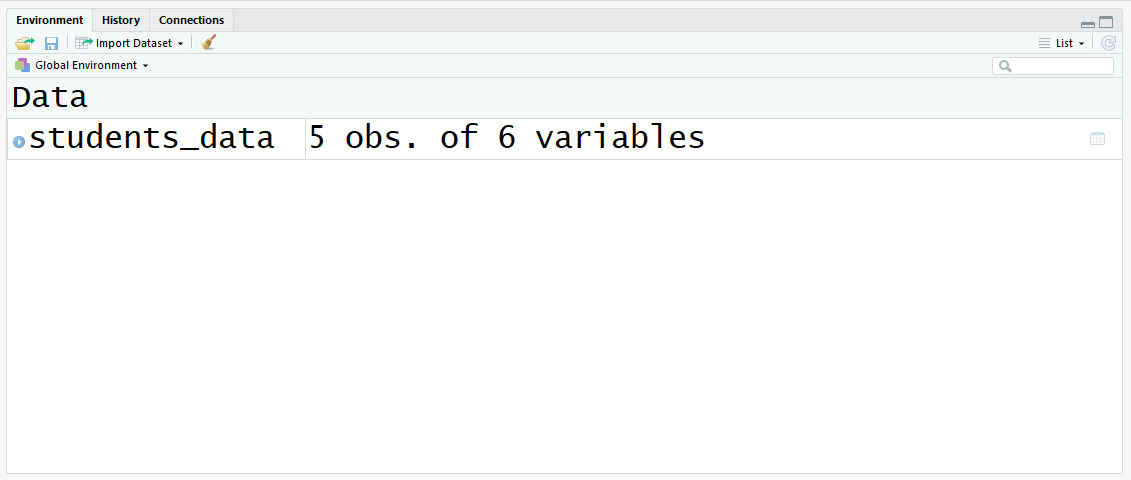
Now y'all have a variable defined in the environs! Let's run into what data frames are and how they are closely related to CSV files.
🔸 Introduction to Information Frames
Information frames are the standard digital format used to shop statistical information in the form of a tabular array. When you read a CSV file in R, a data frame is generated.
Nosotros tin confirm this past checking the type of the variable with the class office:
> form(students_data) [1] "information.frame" Information technology makes sense, right? CSV files contain data represented in the form of a table and data frames represent that tabular data in your lawmaking, so they are deeply connected.
If yous enter this variable in the interactive panel, you will run across the content of the CSV file:
> students_data first_name last_name age num_siblings num_pets eye_color 1 Emily Dawson xv 2 5 Blueish ii Rose Patterson 14 5 0 Light-green iii Alexander Smith 16 0 2 Brown 4 Nora Navona 16 iv x GREEN v Gino Sand 17 3 8 Blue More Data About the Data Frame
Yous have several unlike alternatives to come across the number of variables and observations of the information frame:
- Your get-go option is to wait at the elevation right panel that shows the variables that are currently divers in the environment. This data frame has 5 observations (rows) and six variables (columns):
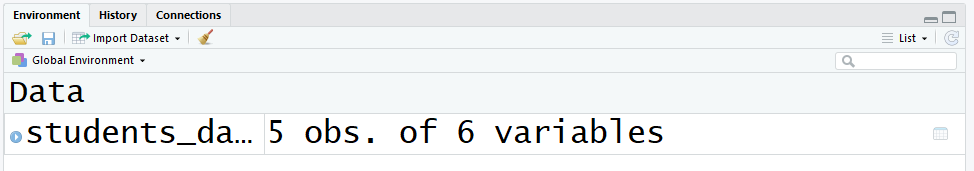
- Another alternative is to utilise the functions
nrowandncolin the interactive console or in your program, passing the data frame as statement. We go the same results: v rows and 6 columns.
> nrow(students_data) [ane] 5 > ncol(students_data) [one] 6 - You lot can also run across more data about the data frame using the
strfunction:
> str(students_data) 'data.frame': 5 obs. of vi variables: $ first_name : Factor due west/ 5 levels "Alexander","Emily",..: 2 5 1 4 three $ last_name : Factor due west/ 5 levels "Dawson","Navona",..: 1 three 5 2 iv $ age : int xv 14 16 16 17 $ num_siblings: int 2 five 0 4 3 $ num_pets : int five 0 ii 10 8 $ eye_color : Factor due west/ 3 levels "Bluish","Brownish",..: 1 3 2 3 1 This function (applied to a data frame) tells yous:
- The number of observations (rows).
- The number of variables (columns).
- The names of the variables.
- The information types of the variables.
- More information about the variables.
You tin can see that this function is really great when y'all want to know more about the data that you are working with.
💡 Tip: In R, a "Gene" is a qualitative variable, which is a variable whose values represent categories. For example, eye_color has the values "BLUE", "BROWN", "Light-green" which are categories, so as you lot tin can meet in the output of str above, this variable is automatically defined equally a "cistron" when the CSV file is read in R.
🔹 Data Frames: Key Operations and Functions
Now yous know how to see more information about the data frame. But the magic of information frames lies in the amazing capabilities and functionality that they offer, so let's come across this in more than detail.
How to Access A Value of a Information Frame
Information frames are similar matrices, then you can admission individual values using two indices surrounded by foursquare brackets and separated past a comma to indicate which rows and which columns you would similar to include in the result, like this:

For example, if we want to access the value of eye_color (column 6) of the fourth student in the data (row 4):
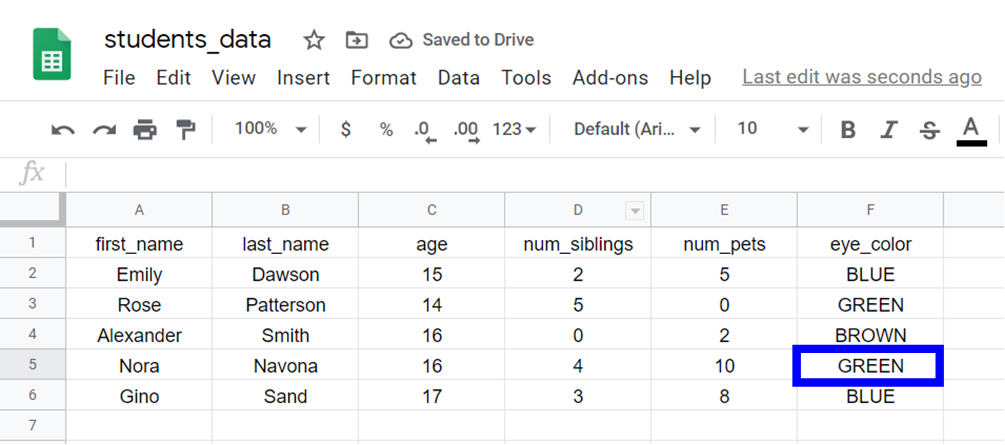
Nosotros demand to use this command:
> students_data[four, six] 💡 Tip: In R, indices first at 1 and the first row with the names of the variables is not counted.
This is the output:
[i] Dark-green Levels: BLUE BROWN Dark-green You tin meet that the value is "GREEN". Variables of type "cistron" accept "levels" that represent the dissimilar categories or values that they can take. This output tells us the levels of the variable eye_color.
How to Admission Rows and Columns of a Data Frame
We can also use this syntax to access a range of rows and columns to get a portion of the original matrix, similar this:
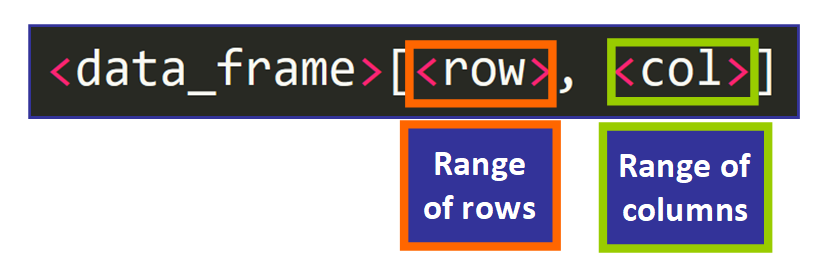

For example, if we want to get the age and number of siblings of the third, fourth, and 5th student in the listing, we would use:
> students_data[3:5, three:4] age num_siblings 3 16 0 iv 16 iv 5 17 3 💡 Tip: The basic syntax to define an interval in R is <outset>:<end>. Note that these indices are inclusive, then the third and fifth elements are included in the case above when we write 3:5.
If we desire to get all the rows or columns, nosotros merely omit the interval and include the comma, like this:
> students_data[3:5,] first_name last_name historic period num_siblings num_pets eye_color iii Alexander Smith 16 0 two Brown iv Nora Navona 16 four 10 Green 5 Gino Sand 17 3 eight Blue We did non include an interval for the columns subsequently the comma in students_data[3:5,], so nosotros become all the columns of the data frame for the iii rows that nosotros specified.
Similarly, nosotros can get all the rows for a specific range of columns if we omit the rows:
> students_data[, 1:3] first_name last_name historic period 1 Emily Dawson 15 2 Rose Patterson xiv three Alexander Smith sixteen 4 Nora Navona xvi 5 Gino Sand 17 💡 Tip: Find that you still need to include the comma in both cases.
How to Access a Column
There are three ways to access an entire cavalcade:
- Option #1: to access a column and return information technology as a information frame, you can use this syntax:

For example:
> students_data["first_name"] first_name 1 Emily 2 Rose 3 Alexander 4 Nora five Gino - Option #ii: to get a column as a vector (sequence), you can use this syntax:

💡 Tip: Notice the use of the $ symbol.
For example:
> students_data$first_name [i] Emily Rose Alexander Nora Gino Levels: Alexander Emily Gino Nora Rose - Option #3: You lot can likewise utilise this syntax to get the column equally a vector (come across below). This is equivalent to the previous syntax:
> students_data[["first_name"]] [1] Emily Rose Alexander Nora Gino Levels: Alexander Emily Gino Nora Rose How to Filter Rows of a Information Frame
You can filter the rows of a information frame to get a portion of the matrix that meets sure conditions.
For this, we use this syntax, passing the condition as the outset element inside square brackets, then a comma, and finally leaving the second element empty.

For example, to get all rows for which students_data$historic period > 16, we would use:
> students_data[students_data$age > xvi,] first_name last_name age num_siblings num_pets eye_color 5 Gino Sand 17 3 8 BLUE We get a data frame with the rows that meet this condition.
Filter Rows and Choose Columns
You tin combine this status with a range of columns:
> students_data[students_data$age > 16, 3:6] age num_siblings num_pets eye_color v 17 three viii Bluish Nosotros get the rows that meet the status and the columns in the range 3:half-dozen.
🔸 How to Modify Data Frames
You tin alter individual values of a data frame, add columns, add rows, and remove them. Let'south see how you tin can practise this!
How to Modify A Value
To modify an private value of the information frame, you need to use this syntax:
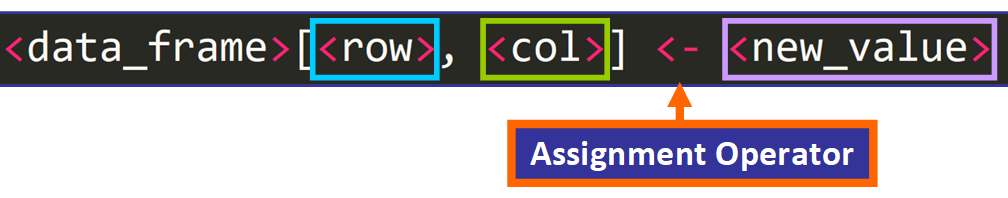
For instance, if we desire to alter the value that is currently at row 4 and column vi, denoted in blue right here:

Nosotros need to employ this line of lawmaking:
students_data[4, 6] <- "BROWN" 💡 Tip: You can also use = equally the assignment operator.
This is the output. The value was changed successfully.
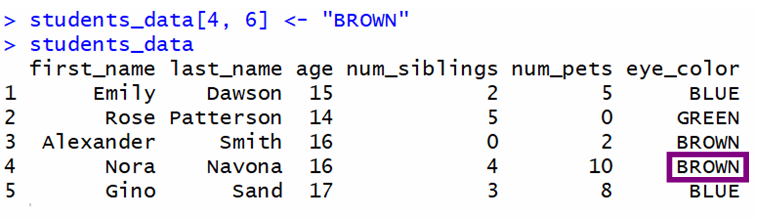

💡 Tip: Remember that the first row of the CSV file is not counted as the showtime row because it has the names of the variables.
How to Add together Rows to a Data Frame
To add a row to a data frame, you need to employ the rbind part:

This role takes two arguments:
- The information frame that you desire to modify.
- A list with the data of the new row. To create the listing, you lot tin can utilize the
list()function with each value separated by a comma.
This is an example:
> rbind(students_data, listing("William", "Smith", 14, 7, 3, "BROWN")) The output is:
first_name last_name age num_siblings num_pets eye_color one Emily Dawson xv 2 5 Blue two Rose Patterson 14 5 0 Greenish iii Alexander Smith 16 0 2 BROWN 4 Nora Navona sixteen four 10 BROWN 5 Gino Sand 17 iii 8 BLUE half-dozen <NA> Smith 14 7 3 BROWN Merely look! A warning message was displayed:
Alert message: In `[<-.factor`(`*tmp*`, ri, value = "William") : invalid gene level, NA generated And notice the first value of the sixth row, it is <NA>:
six <NA> Smith 14 7 3 Dark-brown This occurred because the variable first_name was defined automatically as a factor when nosotros read the CSV file and factors have fixed "categories" (levels).
You cannot add a new level (value - "William") to this variable unless you read the CSV file with the value Imitation for the parameter stringsAsFactors, as shown below:
> students_data <- read.csv("students_data.csv", stringsAsFactors = FALSE) 
Now, if nosotros endeavor to add together this row, the data frame is modified successfully.
> students_data <- rbind(students_data, list("William", "Smith", 14, 7, 3, "Chocolate-brown")) > students_data first_name last_name age num_siblings num_pets eye_color 1 Emily Dawson 15 2 5 Blue 2 Rose Patterson xiv v 0 GREEN 3 Alexander Smith 16 0 2 Brownish 4 Nora Navona 16 4 10 GREEN 5 Gino Sand 17 3 viii Bluish 6 William Smith xiv 7 iii Dark-brown 💡 Tip: Note that if you lot read the CSV file again and assign it to the same variable, all the changes made previously will be removed and you will see the original information frame. You demand to add this argument to the first line of code that reads the CSV file and then make changes to it.
How to Add Columns to a Information Frame
Calculation columns to a data frame is much simpler. Yous need to use this syntax:
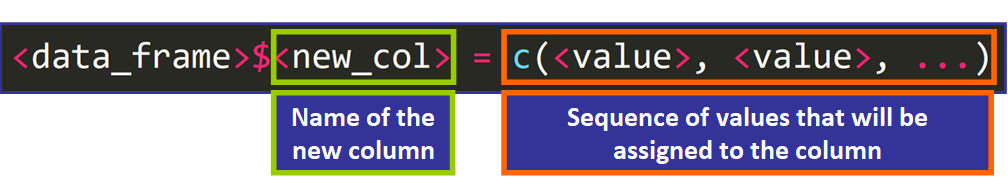
For instance:
> students_data$GPA <- c(4.0, 3.v, 3.2, three.15, 2.9, iii.0) 💡 Tip: The number of elements has to be equal to the number of rows of the data frame.
The output shows the data frame with the new GPA column:
> students_data first_name last_name age num_siblings num_pets eye_color GPA 1 Emily Dawson xv two 5 Blueish 4.00 2 Rose Patterson fourteen 5 0 GREEN 3.fifty iii Alexander Smith 16 0 2 Dark-brown three.20 4 Nora Navona 16 iv 10 Green 3.15 5 Gino Sand 17 3 8 BLUE two.90 6 William Smith 14 seven 3 Dark-brown 3.00 How to Remove Columns
To remove columns from a data frame, yous need to use this syntax:

When yous assign the value Zippo to a column, that cavalcade is removed from the data frame automatically.
For example, to remove the age column, we utilize:
> students_data$historic period <- NULL The output is:
> students_data first_name last_name num_siblings num_pets eye_color GPA 1 Emily Dawson two 5 BLUE iv.00 2 Rose Patterson 5 0 GREEN 3.fifty 3 Alexander Smith 0 2 Chocolate-brown three.20 4 Nora Navona four 10 Light-green 3.15 5 Gino Sand 3 viii Blueish 2.90 half-dozen William Smith 7 three BROWN 3.00 How to Remove Rows
To remove rows from a data frame, yous tin use indices and ranges. For example, to remove the first row of a information frame:

The [-1,] takes a portion of the data frame that doesn't include the first row. Then, this portion is assigned to the same variable.
If we take this data frame and we want to delete the first row:

The output is a data frame that doesn't include the start row:
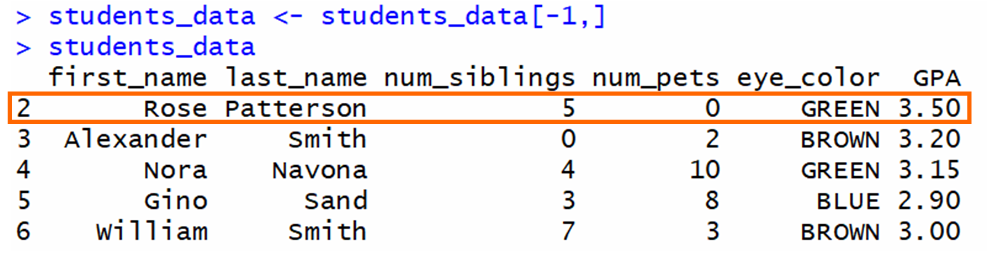
In full general, to remove a specific row, you need to utilise this syntax where <row_num> is the row that you want to remove:

💡 Tip: Notice the - sign before the row number.
For example, if we want to remove row iv from this information frame:

The output is:
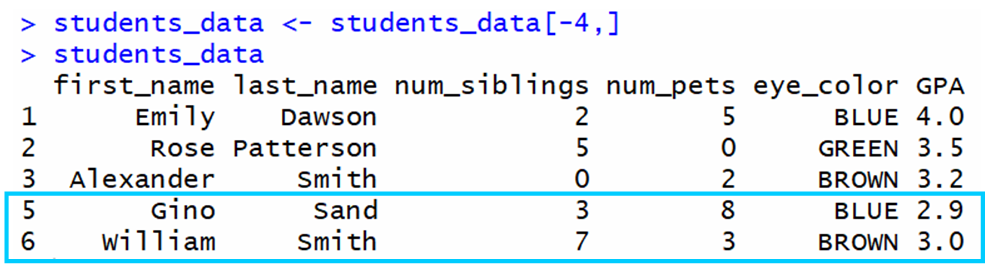
As you can see, row 4 was successfully removed.
🔹 In Summary
- CSV files are Comma-Separated Values Files used to represent data in the form of a table. These files tin can be read using R and RStudio.
- Data frames are used in R to represent tabular information. When you read a CSV file, a data frame is created to store the information.
- You can access and modify the values, rows, and columns of a data frame.
I really hope that yous liked my article and plant it helpful. At present you can piece of work with data frames and CSV files in R.
If you lot liked this article, consider enrolling in my new online course "Introduction to Statistics in R - A Practical Arroyo "
Acquire to code for gratuitous. freeCodeCamp's open source curriculum has helped more than than 40,000 people get jobs as developers. Get started
Source: https://www.freecodecamp.org/news/how-to-work-with-data-frames-and-csv-files-in-r/
0 Response to "How to Read a Column From Csv File in R"
Post a Comment